The UNC Vironomics (viral genomics) Core aims to facilitate ZIKV research projects at UNC, from translational research, to animal studies, to high-throughput screening.
Zika virus (ZIKV) is a newly emerging disease in the Americas. Most individuals who are infected with ZIKV are asymptomatic, and non-pregnant symptomatic patients generally present with mild flu-like illness. Unfortunately, ZIKV infection may be linked to profound damage to the developing fetus including microcephaly and miscarriage.
The CDC recommends laboratory testing for patients presenting with flu-like symptoms who have traveled to areas where ZIKV is circulating, and the CDC now recommends that blood banks test for ZIKV in donated blood. Despite this wide-spread need for ZIKV testing, there are no currently FDA approved nucleic acid diagnostic tests for ZIKV.
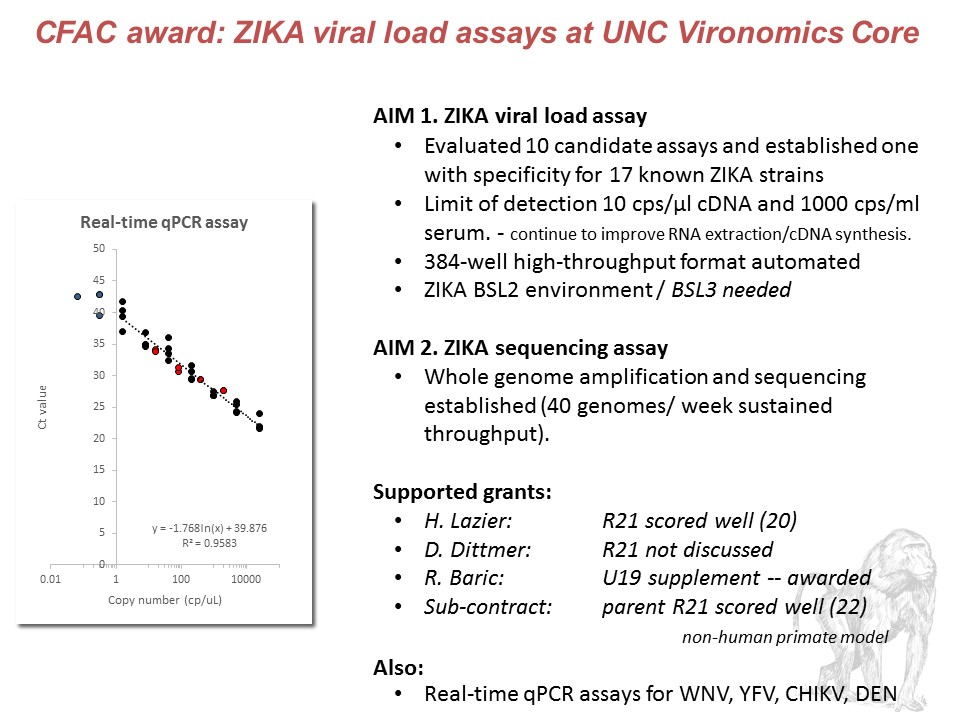
With funding from the Core Facilities Advocacy Committee (CFAC), The UNC Vironomics Core has successfully developed a high-throughput RT-qPCR viral load assay for ZIKV. After evaluation of 10 published assays for ZIKV nucleic acid detection, the Vironomics Core has developed an improved ZIKV viral load assay. The Vironomics Core continues to improve the sensitivity of this assay, as well as evaluating a whole genome sequencing assay compatible with sequencing virus from clinical samples. Additionally, the Vironomics Core continues to develop high-throughput RT-qPCR assays for flaviviruses commonly found in regions with ZIKV circulation: West Nile, Yellow Fever, Dengue, and Chikungunya.
There is an immediate need for increased research about ZIKV biology, the potential for fetal pathology, and disease prevention. In response to the immediate need for more understanding about ZIKV biology and pathogenesis, the NIH has called for priority funding towards ZIKV research (NOT-AI-16-026, NOT-HD-16-004). The Vironomics Core aims to provide robust ZIKV viral load services in order to facilitate the UNC research community in new studies of this important emerging pathogen.