Paper titled “Site-Specific Silencing of Regulatory Elements as a Mechanism of X Inactivation.”
Mauro Calabrese, a postdoctoral fellow in Terry Magnuson’s lab, is first-author of an article in Cell entitled “Site-Specific Silencing of Regulatory Elements as a Mechanism of X Inactivation.” Coauthors with current or previous Department of Genetics affiliations are Wei Sun, Joshua Mugford, Lucy Williams, Della Yee, Joshua Starmer, and Piotr Mieczkowski, with Terry as senior author.
Summary
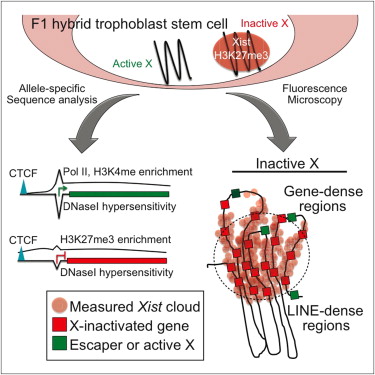
“The inactive X chromosome’s (Xi) physical territory is microscopically devoid of transcriptional hallmarks and enriched in silencing-associated modifications. How these microscopic signatures relate to specific Xi sequences is unknown. Therefore, we profiled Xi gene expression and chromatin states at high resolution via allele-specific sequencing in mouse trophoblast stem cells. Most notably, X-inactivated transcription start sites harbored distinct epigenetic signatures relative to surrounding Xi DNA. These sites displayed H3-lysine27-trimethylation enrichment and DNaseI hypersensitivity, similar to autosomal Polycomb targets, yet excluded Pol II and other transcriptional hallmarks, similar to nontranscribed genes. CTCF bound X-inactivated and escaping genes, irrespective of measured chromatin boundaries. Escape from X inactivation occurred within, and X inactivation was maintained exterior to, the area encompassed by Xist in cells subject to imprinted and random X inactivation. The data support a model whereby inactivation of specific regulatory elements, rather than a simple chromosome-wide separation from transcription machinery, governs gene silencing over the Xi.”
Highlights
“► RNA-Seq analysis finds 13% of genes escape X inactivation in trophoblast stem cells ► Inactive X regulatory elements lack Pol II binding yet retain DNaseI hypersensitivity ► CTCF binds across the inactive X, mostly to locations shared with the active X ► X-inactivated genes near escapers are often separated from the measured Xist cloud.”