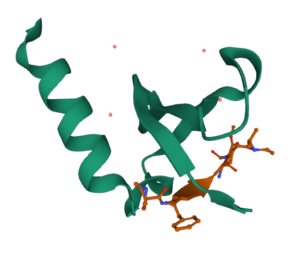
The structure of a peptidomimetic ligand UNC5246 bound to the chromodomain of MPP8 (PDB ID 7M5U) was solved by McGinty lab in collaboration with Lindsay Ingerman-James group and published in ACS Chemical Biology. In this paper, a biotinylated version of the ligand was used as an affinity reagent to identify novel MPP8 interactors.



Abstract
The interpretation of histone post-translational modifications (PTMs), specifically lysine methylation, by specific classes of “reader” proteins marks an important aspect of epigenetic control of gene expression. Methyl-lysine (Kme) readers often regulate gene expression patterns through the recognition of a specific Kme PTM while participating in or recruiting large protein complexes that contain enzymatic or chromatin remodeling activity. Understanding the composition of these Kme-reader-containing protein complexes can serve to further our understanding of the biological roles of Kme readers, while small molecule chemical tools can be valuable reagents in interrogating novel protein–protein interactions. Here, we describe our efforts to target the chromodomain of M-phase phosphoprotein 8 (MPP8), a member of the human silencing hub (HUSH) complex and a histone 3 lysine 9 trimethyl (H3K9me3) reader that is vital for heterochromatin formation and has specific roles in cancer metastasis. Utilizing a one-bead, one-compound (OBOC) combinatorial screening approach, we identified UNC5246, a peptidomimetic ligand capable of interacting with the MPP8 chromodomain in the context of the HUSH complex. Additionally, a biotinylated derivative of UNC5246 facilitated chemoproteomics studies which revealed hepatoma-derived growth factor-related protein 2 (HRP2) as a novel protein associated with MPP8. HRP2 was further shown to colocalize with MPP8 at the E-cadherin gene locus, suggesting a possible role in cancer cell plasticity.
Link to paper
A Peptidomimetic Ligand Targeting the Chromodomain of MPP8 Reveals HRP2’s Association with the HUSH Complex
DOI: 10.1021/acschembio.1c00429
Link to structure
Structure of peptidomimetic ligand bound to MPP8 chromodomain in the RCSB Protein Data Bank.
About McGinty lab
McGinty lab’s research focus is on the nucleosome is a dynamic epigenetic signaling hub for genome-templated processes. We use high-resolution structural biology, protein chemistry, and proteomics to understand the molecular mechanisms that govern epigenetic signaling in health and disease. Continue reading more about their work on the McGinty lab website.
Read Twitter threads from September 1, 2021.
There are 250 new #PDB entries this week…
and

