 Conserved helix-turn-helix motif contributes differentially to #polyubiquitination in various #cellcycle E2 enzymes – via distinct donor/acceptor #ubiquitin interactions
Conserved helix-turn-helix motif contributes differentially to #polyubiquitination in various #cellcycle E2 enzymes – via distinct donor/acceptor #ubiquitin interactions
Abstract:
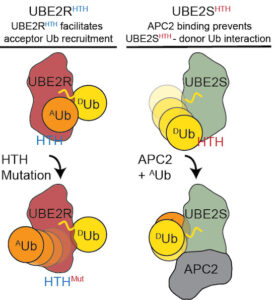
Polyubiquitination by E2 and E3 enzymes is crucial to cell cycle control, epigenetic regulation, and development. The hallmark of the E2 family is the ubiquitin (Ub)-conjugating (UBC) domain that forms a dynamic thioester conjugate with ubiquitin (E2~Ub). Numerous studies have focused on E2 surfaces, such as the N-terminal and crossover helices, that directly interact with an E3 or the conjugated ubiquitin to stabilize the active, “closed” state of the E2~Ub. However, it remains unclear how other E2 surfaces regulate ubiquitin transfer. Here, we demonstrate the helix–turn–helix (HTH) motif of the UBC tunes the intrinsic polyubiquitination activity through distinct functions in different E2s. Interestingly, the E2HTH motif is repurposed in UBE2S and UBE2R2 to interact with the conjugated or acceptor ubiquitin, respectively, modulating ubiquitin transfer. Furthermore, we propose that Anaphase-Promoting Complex/Cyclosome binding to the UBE2SHTHreduces the conformational space of the flexible E2~Ub, demonstrating an atypical E3-dependent activation mechanism. Altogether, we postulate the E2HTH motif evolved to provide new functionalities that can be harnessed by E3s and permits additional regulation to facilitate specific E2-E3-mediated polyubiquitination.
Link to publication:
A collaborative effort led by Kaeli Welsh (Pharmacology) who is the lead author amongst the colleagues, Nick Brown (Pharmacology) and Joe Harrison (a former postdoc of Brian Kuhlman) and in BCBP (Josh Boyer, Brenda Temple, Derek Bolhuis, Qi Zhang) and other colleagues from the University of the Pacific: Chemistry & Mechanical Engineering departments, Department of Cell Biology, Blavatnik Institute of Harvard Medical School.